
- Home
- CliniDeID
CliniDeID
Automatic clinical text de-identification
CliniDeID® automatically de-identifies clinical notes and structured data according to the HIPAA Safe Harbor method. It accurately finds identifiers and tags or replaces them with realistic surrogates for better anonymity. It improves access to richer, more detailed, and more accurate clinical data for clinical researchers. It eases research data sharing, and helps healthcare organizations protect patient data confidentiality.
Available as free open source software (GPL v3 license): download link.
Concerned about exposing patients to privacy breaches?
About HIPAA violations?
Trying to share clinical data for research?


- Increase patient trust through the ethical protection of confidentiality.
- Magnify the impact of clinical data for research by providing the ability to reuse data without patient informed consent.
- Greatly reduce financial risks by avoiding fines and other penalties that could otherwise arise from a leak of patient data.
- Save money by reducing the cost and increasing the efficiency of clinical data de-identification.
- Expand and scale existing research opportunities by providing richer, more detailed clinical data.
- Uncover and facilitate new research opportunities through shareable, de-identified clinical data as well as provide larger, NIH-sponsored research with required data sharing capabilities.
- “Future-proof” clinical data for unforeseen research opportunities by allowing records to be accurately linked even after de-identification.

Features
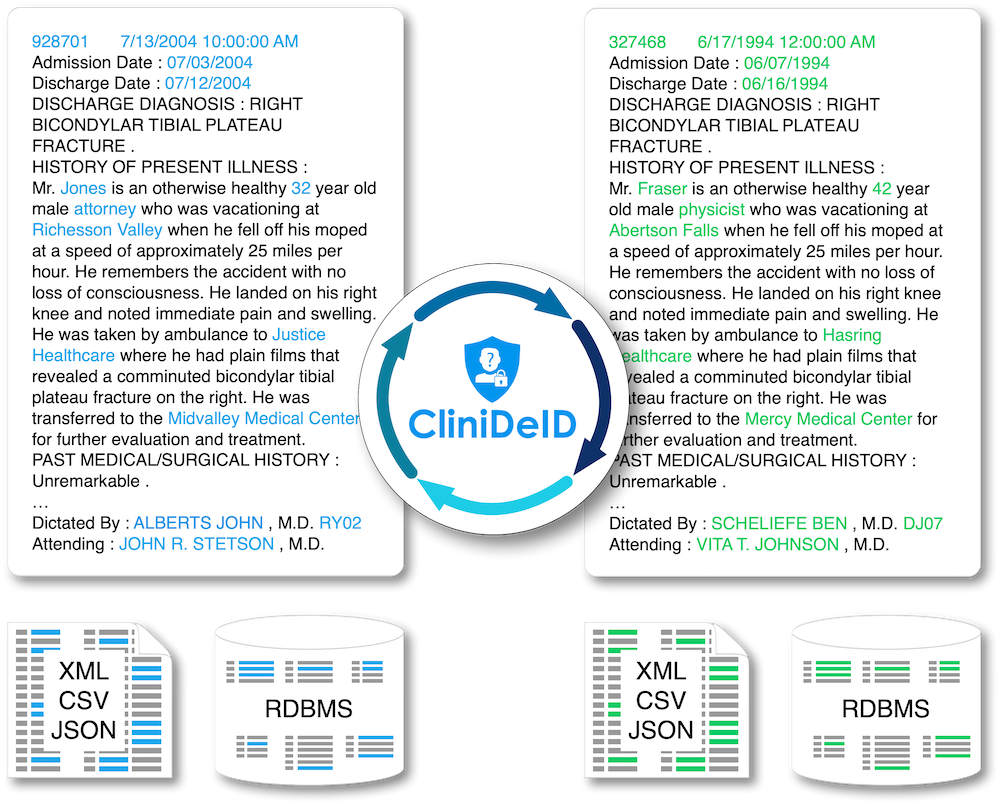
Clinical text de-identification: Uses advanced artificial intelligence algorithms to accurately identify all mentions of identifiers (PII) in unstructured clinical text notes and replaces them with realistic surrogates (PII resynthesis) or tags, as desired. Does not rely on any known identifiers but can use known identifiers to double-check the PII identification if available. Generalizes well to all common types of clinical notes.
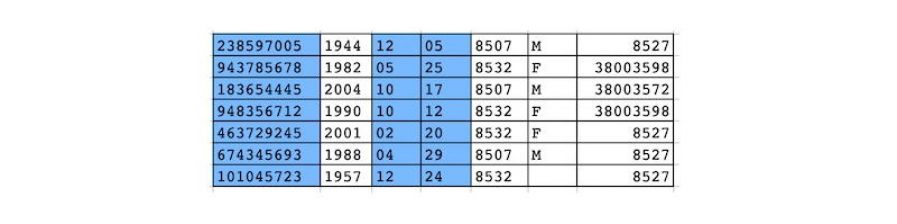
Structured data de-identification, integrated with unstructured text de-identification for consistent de-identification throughout the patient record (in CliniDeiD-Complete). Currently compatible with standard data models: OMOP CDM v5.3 and v6; HL7 FHIR coming soon)

Replacement of identified PII with realistic surrogates and consistently across the whole patient record (PII resynthesis), or with tags (generic or PII categories).

Highly accurate identification of PII (as demonstrated in several peer-reviewed evaluations and comparisons available at the bottom of this page)

Multiple input and output data formats: plain text, HL7 CDA , relational databases (PostgreSQL, Oracle , MySQL, MS SQL Server, DB2)





